The surface of a cell is a dynamic place - like children in a busy playground, molecules move around in apparently random patterns. However, there is order in this ostensibly chaotic system. Since many of the cell's sensors reside on its surface, they often aggregate specifically in response to a stimulus to transmit information about the environment into the cell. But what are the rules that govern the movement of such molecules on the cell surface? One way to answer this question is to model these processes by reconstructing the cell surface from scratch. Now, scientists from the National Centre for Biological Sciences (NCBS) in Bangalore have managed to do exactly that - construct the cell surface from its constituent parts, namely, a mixture of lipids and proteins. This reconstruction creates a crucial tool researchers can use to test theories on cell surface dynamics.
A cell's surface is covered by a cell membrane, traditionally described as a lipid bilayer embedded with proteins. This fairly simplistic model also assumes that molecules in the cell membrane move freely in a 2-dimensional space - one could imagine this as a 'sea' of lipids with protein 'ships' moving freely on its surface. However, in the light of new observations, molecular movements on the cell surface are now known to be non-random, incredibly complex, and do not seem to follow simple thermodynamic rules. The 'active composite model' of the cell surface is one of the latest theories that attempts to explain such molecular movements. This theory was developed by a close collaboration between experimental biologist Prof. Satyajit Mayor at NCBS, theoretical physicist Prof. Madan Rao at NCBS and RRI (Raman Research Institute) and colleagues from their respective groups.
Recently, these two teams came together again with scientists from the University of California San Francisco (UCSF) to create an experimental system that would allow them to test predictions made by the active composite model. This experimental system would be a minimal model of the cell surface constructed from its basic components - purified lipids and proteins known to be part of the cell surface. The active composite model visualises the cell surface as not just the cell membrane, but as an amalgamation of two elements - the cell membrane and an interwoven mesh of the protein 'actin' that forms a thin layer or 'actin cortex' just below the cell membrane. Another protein, 'myosin' that interacts with actin, is a molecular motor that causes movement in the actin meshwork when supplied with energy. Many of the cell surface proteins whose movements have baffled scientists are known to be linked directly or indirectly to the dynamic actin meshwork that lies just below the cell membrane.
"This is just a beginning but an important one," says Prof Satyajit Mayor. "Important because it allows one to test ideas that have come from theory built around providing an explanation for active organization at the surface of a living cell. It's an exciting beginning since the feasibility of this simple minimal system opens up huge possibilities to explore the world of a living cell in a test tube system where every element is under our control. This work is inspired by the adage 'what we understand we should be able to build' and this is in trying to understand the principles behind how a living material, the cell surface, works," he adds.
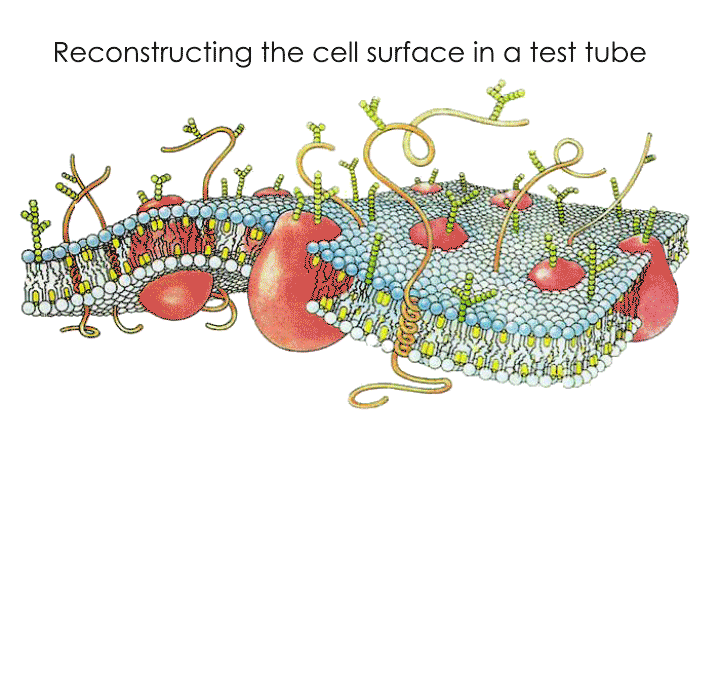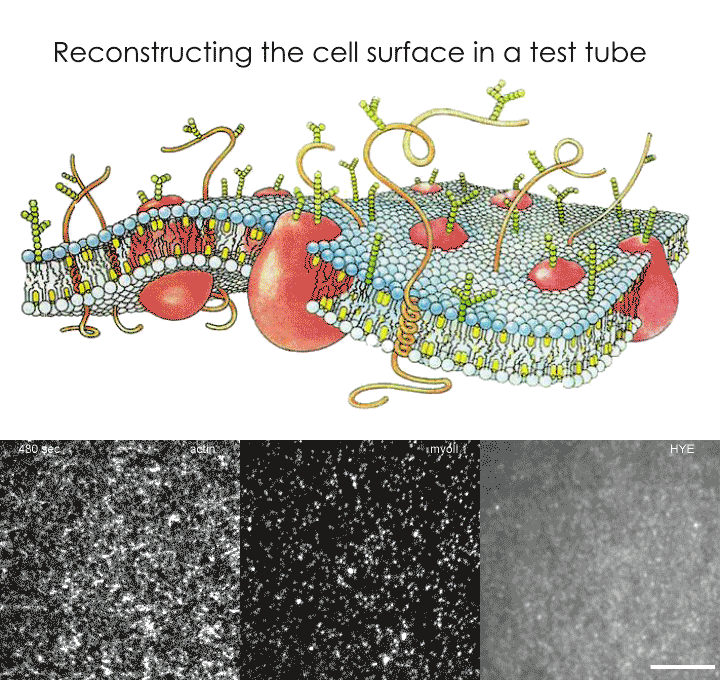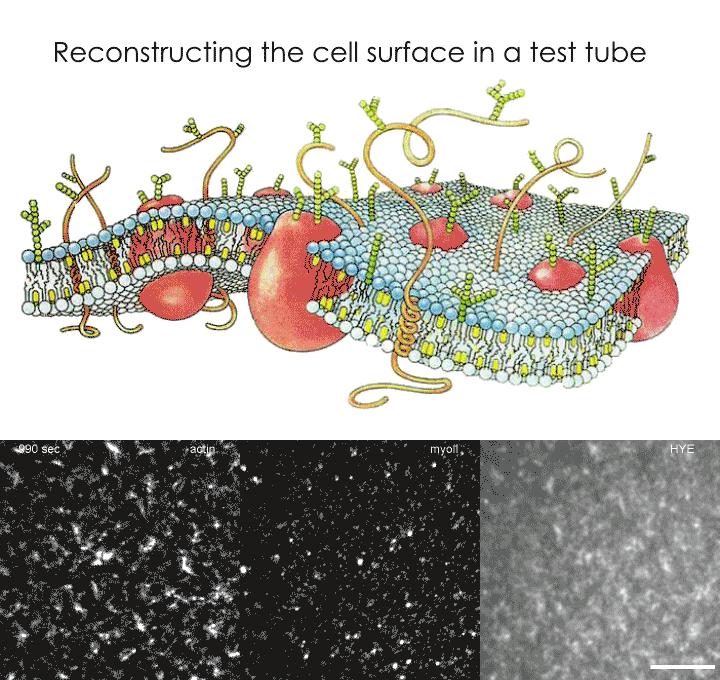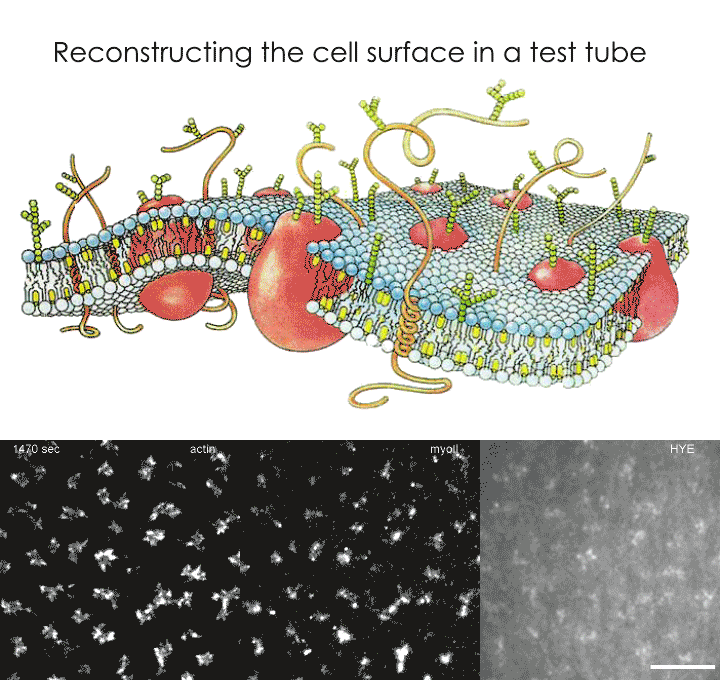
As proposed by the active composite model, the researchers decided to recreate a cell surface as an assembly of a lipid-based membrane and an actin meshwork. This artificial cell surface was therefore constructed using a lipid bilayer, actin and a fluorescent protein specially designed to be associated with the membrane while also being linked to actin. Using various microscopic techniques, the group were able to study the behaviour of the construct via the patterns formed by the fluorescent proteins. True to the prediction of the 'active composite model', the movement of these actin-bound proteins were found to be dependent on the dynamics of the actin meshwork. When the molecular motor, myosin, was added and energy in the form of ATP provided, the forces generated by actin-myosin interactions drove the movements of these proteins. The bulk motion of the actin meshwork, specifically during contractile flows, caused these proteins to aggregate through a process known as advection.
Once the energy from ATP was exhausted, and the system reached equilibrium, the actin-bound proteins aggregated to form distinct bundle or aster-like structures based on the component characteristics of the system. The shape and size of these aggregations were dependent on the concentration of actin, length of actin filaments and the ratio of actin to myosin motors. Bundles were formed in conditions with high actin concentration, long actin filaments and high actin-to-myosin ratios; whereas polar asters were formed under condition with low actin concentrations, short actin filaments and low actin-to-myosin ratios. A further continuous influx of ATP to such equilibrium conditions caused the bundles and asters to dissolve, reform as localised accumulations and dissolve again in close mimicry of the highly dynamic cortex of a living cell. This continuous remodelling of the actin meshwork also led to 'giant number fluctuations', which are large spatial and temporal variations in actin-bound fluorescent protein densities. This phenomenon is also observed in Glycophosphatidylinositol (GPI) anchored proteins on living cells, indicating that the reconstructed minimal model is likely to be a good representation of the surface of a live cell.
"The importance of active or energy consuming processes in understanding biological phenomena is becoming more and more evident. This is an emerging field in biology called 'active mechanics'. Often, the emerging organisation of biological molecules are not clear, and theoretical explanations for such observations are also far from complete," says Darius Köster, the lead author of the study that was published in the leading journal PNAS (Proceedings of the National Academy of Sciences of the United States of America). "This makes it important to have proper experimental tools that go hand in hand with theory to test and improve our understanding of such systems. Our current study describes the creation of an experimental system that will serve us in this," he adds.
"The motivation behind this work is to analyse mechanisms influencing the dynamics and organisation of molecules on the cell surface," says Kabir Husain, another author in this study.
Processes like cell growth, division, immune recognition and many others are dependent on the organisation of protein receptors and other associated molecules on the cell surface. This implies that the ability of the cell to reliably control the organisation of its surface molecules is crucial to its survival and function. With the recreation of the cell surface in a test tube, scientists have gained a solid experimental footing in the race to comprehend the mechanics of cell surface organisation.
This work has been published in a paper titled "Actomyosin dynamics drive local membrane component organization in an invitro active composite layer" in Feb 2016 in the journal PNAS. The work has been supported by grant RGP0027/2012 from the HFSPO, a DST-SERB JC Bose Fellowship and intramural support from NCBS-TIFR to Satyajit Mayor and an AXA Research Fellowship award to Dr Darius Koster. Prof Madan Rao is a member of the Simons Centre for the Study of Living Machines at NCBS.









0 Comments