Researchers from Bangalore and Dresden discover a unique two-component molecular motor that uses a kind of renewable chemical energy to pull vesicles toward membrane-bound organelles.
Cells have a fascinating feature to neatly organize their interior by using tiny protein machines called molecular motors that generate directed movements. Most of them use a common type of fuel, a kind of chemical energy, called ATP to operate. Now researchers at the Simons Centre for the Study of Living Machines at the National Centre for Biological Sciences (NCBS), the Max Planck Institute of Molecular Cell Biology and Genetics (MPI-CBG), the Cluster of Excellence Physics of Life (PoL) and the Biotechnology Center (BIOTEC) of the Technische Universität Dresden (TUD) in Dresden, Germany, have discovered a novel molecular system that uses an alternative chemical energy and employs a novel mechanism to perform mechanical work. By repeatedly contracting and expanding, this molecular motor functions similarly to a classical Stirling engine and helps to distribute cargo to membrane-bound organelles. It is the first motor using two components, two differently sized proteins, Rab5 and EEA1, and is driven by GTP instead of ATP. The results are published in the journal Nature Physics.
Motor proteins are remarkable molecular machines within a cell that convert chemical energy, stored in a molecule called ATP, into mechanical work. The most prominent example is myosin which helps our muscles to move. In contrast, GTPases which are small proteins have not been viewed as molecular force generators. One example is a molecular motor composed of two proteins, EEA1 and Rab5. In 2016, an interdisciplinary team of cell biologists and biophysicists at the MPI-CBG discovered (in a result published in Nature) that the small GTPase protein Rab5 could trigger a contraction in EEA1. These string-shaped tether proteins can recognize the Rab5 protein present in a vesicle membrane and bind to it. The binding of the much smaller Rab5 sends a message along the elongated structure of EEA1, thereby increasing its flexibility, similar to how cooking softens spaghetti. Such flexibility change produces a force that pulls the vesicle towards the target membrane, where docking and fusion occur. However, the team also hypothesized that EEA1 could switch between a flexible and a rigid state, similar to a mechanical motor motion, simply by interacting with Rab5 alone.
This is where the current research sets in, taking shape via the doctoral work of the two first authors of the study. Anupam Singh from the group of Shashi Thutupalli, a biophysicist at the Simons Centre for the Study of Living Machines at the NCBS and Joan Antoni Soler from Marino Zerial's research group at MPI-CBG set out to experimentally observe this motor in action.
With an experimental design to investigate the dynamics of the EEA1 protein in mind, Anupam Singh spent three months at the MPI-CBG in 2019. "When I met Joan, I explained to him the idea of measuring the protein dynamics of EEA1. But these experiments required specific modifications to the protein that allowed measurement of its flexibility based on its structural changes,” says Anupam. Joan Antoni Soler's expertise in protein biochemistry was a perfect fit for this challenging task. "I was delighted to learn that the approach to characterize the EEA1 protein could answer whether EEA1 and Rab5 form a two-component motor, as previously suspected. I realized that the difficulties in obtaining the correct molecules could be solved by modifying the EEA1 protein to allow fluorophores to attach to specific protein regions. This modification would make it easier to characterize the protein structure and the changes that can occur when it interacts with Rab5," explains Joan Antoni.
Armed with the suitable protein molecules and the valuable support of co-author Janelle Lauer, a senior postdoctoral researcher in Marino Zerial's research group, Joan and Anupam were able characterize the dynamics of EEA1 thoroughly using the advanced laser scanning microscopes provided by the light microscopy facilities at the MPI-CBG and the NCBS. Strikingly, they discovered that the EEA1 protein could undergo multiple flexibility transition cycles, from rigid to flexible and back again, driven solely by the chemical energy released by its interaction with the GTPase Rab5. These experiments showed that EEA1 and Rab5 form a GTP-driven two-component motor. To interpret the results, Marcus Jahnel, one of the corresponding authors and research group leader at PoL and BIOTEC, developed a new physical model to describe the coupling between chemical and mechanical steps in the motor cycle. Together with Stephan Grill and Shashi Thutupalli, the biophysicists were also able to calculate the thermodynamic efficiency of the new motor system, which is comparable to that of conventional ATP-driven motor proteins.
"Our results show that the proteins EEA1 and Rab5 work together as a two-component molecular motor system that can transfer chemical energy into mechanical work. As a result, they can play active mechanical roles in membrane trafficking. It is possible that the force-generating molecular motor mechanism may be conserved across other molecules and used by several other cellular compartments," Marino Zerial summarizes the study. Marcus Jahnel adds: "I am delighted that we could finally test the idea of an EEA1-Rab5 motor. It's great to see it confirmed by these new experiments. Most molecular motors use a common type of cellular fuel called ATP. Small GTPases consume another type of fuel, GTP, and have been thought of mainly as signaling molecules. That they can also drive a molecular system to generate forces and move things around puts these abundant molecules in an interesting new light." Stephan Grill is equally excited: "It's a new class of molecular motors! This one doesn't move around like the Kinesin motor that transports cargo along microtubules but performs work while staying in place. It's a bit like the tentacles of an octopus."
"The model we used is inspired by that of the classical Stirling engine cycle. While the traditional Stirling engine generates mechanical work by expanding and compressing gas, the two-component motor described uses proteins as the working substrate, with protein flexibility changes resulting in force generation. As a result, this type of mechanism opens up new possibilities for the development of synthetic protein engines," adds Shashi Thutupalli.
Overall, the authors hope that this new interdisciplinary study could open new research avenues in both molecular cell biology and biophysics.
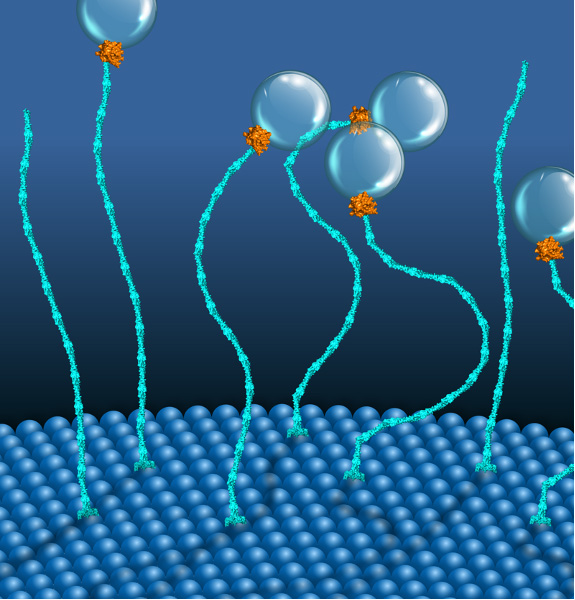
Picture Above: A two-component molecular motor placing vesicles proximal to endosomal membranes. Copyright: MPI-CBG
Original Publication:
Anupam Singh, Joan Antoni Soler, Janelle Lauer, Stephan W Grill, Marcus Jahnel, Marino Zerial, and Shashi Thutupalli: Two-component molecular motor driven by a GTPase cycle, Nature Physics (2023), https://www.nature.com/articles/s41567-023-02009-3
DOI: 10.1038/s41567-023-02009-3
About the National Centre for Biological Sciences (NCBS), Tata Institute of Fundamental Research
The mandate of research at NCBS in Bangalore, India, is the study of biology at all scales. To realize these goals of understanding living systems, researchers at the NCBS approach biological questions from a variety of directions. Particularly, the Simons Center for the Study of Living Machines within the National Centre for Biological Sciences, part of the Tata Institute of Fundamental Research is focused on quantitative and theoretical approaches to understanding living systems. The centre is therefore populated by scientists with a variety of backgrounds in the physical sciences, mathematics, and computer science. The International Centre for Theoretical Sciences (ICTS) of the Tata Institute of Fundamental Research was established in 2007 and is a multi and interdisciplinary centre dedicated to theoretical sciences.
About the MPI-CBG
The Max Planck Institute of Molecular Cell Biology and Genetics (MPI-CBG), located in Dresden, is one of more than 80 institutes of the Max Planck Society, an independent, non-profit organization in Germany. 550 curiosity-driven scientists from over 50 countries ask: How do cells form tissues? The basic research programs of the MPI-CBG span multiple scales of magnitude, from molecular assemblies to organelles, cells, tissues, organs, and organisms. The MPI-CBG invests extensively in Services and Facilities to allow research scientists shared access to sophisticated and expensive technologies.
About the Excellence Cluster Physics of Life (PoL)
Physics of Life (PoL) is one of three Clusters of Excellence at TU Dresden. It focuses on identifying the laws of physics that underlie the organization of life into molecules, cells, and tissues. At the cluster, physicists, biologists, and computer scientists join forces to investigate how active matter organizes itself into predetermined structures in cells and tissues, thus giving rise to life. Pol is funded by the DG as part of the Excellence Strategy. It is a collaboration between scientists from TU Dresden and research institutions of the DRESDEN-concept network, such as the Max Planck Institute of Molecular Cell Biology and Genetics (MPI-CBG), the Max Planck Institute for the Physics of Complex Systems (MPI-PKS), the Leibniz Institute of Polymer Research (IF) and the Helmholtz-Zentrum Dresden-Rossendorf (HZDR).
About the Biotechnology Center of the TU Dresden
The Biotechnology Center (BIOTEC) was founded in 2000 as a central scientific unit of the TU Dresden with the goal of combining modern approaches in molecular and cell biology with the traditionally strong engineering in Dresden. Since 2016, the BIOTEC is part of the central scientific unit "Center for Molecular and Cellular Bioengineering" (CMCB) of the TU Dresden. The BIOTEC is fostering developments in research and teaching within the Molecular Bioengineering research field and combines approaches in cell biology, biophysics and bioinformatics. It plays a central role within the research priority area Health Sciences, Biomedicine and Bioengineering of the TU Dresden.









0 Comments